Total points: 30
Due: Wednesday 25th September, 7pm CEST
Format: IPython Notebook or python program
The number of points in this problem sheet is not directly proportional to the difficulty. In fact, Part 3 is more difficult than Part 1 and 2, but is worth fewer points in terms of the amount of work/code, and are there for people who enjoy challenges. So you can choose to not complete Part 3, and you should still be able to get more than 60% of points from the previous questions if you answer them correctly.
Background¶
This problem set is about DNA data and methods typically applied to this kind of data. As a reminder, DNA is a molecule that encodes genetic instructions for living organisms. DNA is typically found as a double-stranded helix, in which each strand corresponds to a sequence of nucleotides. Each nucleotide consists of a nucleobase (guanine, adenine, thymine, or cytosine) attached to sugars, which are in turn separated from each other by phosphate groups:
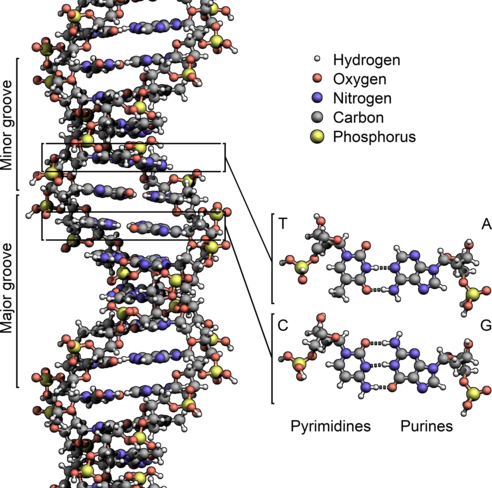
(image from Wikipedia)
The nucleobases are commonly referred to with the letters G (guanine), A (adenine), T (thymine), and C (cytosine). The nucleobases form pairs between the two strands: G pairs with C, and T pairs with A.
DNA is therefore most commonly represented as a sequence of nucleobases, such as GATTACACCTCATTATAAA.
For more information about DNA, see the wikipedia page in German or English.
In this problem set, we will be working with fake genetic data, since real-life data often has added complications, but the functions and techniques uses here are the same or very similar to techniques used on real data.
Part 1 (10 points)¶
Question 1 (6 points)¶
The reverse complement of DNA is found by reversing the DNA sequence, then replacing each base by its complement (A is replaced by T, T is replaced by A, G is replaced by C, and C is replaced by G). For example, the reverse complement of ATGCGGC is GCCGCAT
Write a Python function reverse_complement that takes a DNA sequence as a string, and returns the reverse complement. Test your function by ensuring that reverse_complement('ATGCGGC') is 'GCCGCAT', then find the reverse complement of the following sequence:
ATGCGCGGATCGTACCTAATCGATGGCATTAGCCGAGCCCGATTACGC
Print the result out using print.
Note that this function is NOT needed for the remaining questions below.
Question 2 (4 points)¶
Ribonucleic acid (RNA) is a family of large biological molecules that is transcribed from DNA by the RNA Polymerase enzyme. It consists of a single strand of nucleotides that are identical to the ones found in DNA, with the exception of uracil (U), which replaces thymine (T).
Messenger RNA molecules (or mRNA) are a subset of RNA molecules that are used to pass information from DNA to ribosomes, which then translates the mRNA to protein sequences.
Write a Python function dna_to_mrna that takes a DNA sequence and returns the corresponding mRNA sequence. For example, the DNA sequence ATCGCGAT should produce the mRNA sequence AUCGCGAU (note that you do not need to find the reverse complement of the DNA here)
Part 2 (15 points)¶
Question 3 (8 points)¶
When the mRNA is translated to a protein sequence, each set of three nucleotides, called a codon, is translated into a single amino acid. For example, the codon UUC translates to the amino acid Phenylalanine. Each amino acid can be represented by a single letter - for example Phenylalanine is represented by the letter F. A protein, which is formed from a sequence of amino acids, can therefore be written as a sequence of letters in the same way as DNA or mRNA, but using more of the letters of the alphabet since there are more than four amino acids.
The data/problem_1_codons.txt file contains two columns. The first column gives a list of codons, and the second column gives the corresponding amino acid (represented by a single letter). Certain codons do not correspond to an amino acid, but instead indicate that the amino acid sequence is finished. These are indicated by Stop.
Write a function mrna_to_protein that takes an mRNA sequence (as a string) and returns the sequence of amino acids (as a string), stopping the first time a Stop codon is encountered. Make sure that the file is only read once when running the script (and not every time you want to translate a codon). You will likely need to use a Python dictionary to help.
Finally, write a function dna_to_protein that takes a DNA sequence (as a string) and returns the sequence of amino acids (as a string), making use of the functions that you wrote previously.
Print out the amino acid sequence for the following DNA sequence:
AATCTCTACGGAAGTAGGTCAGTACTGATCGATCAGTCGATCGGGCGGCGATTTCGATCTGATTGTACGGCGGGCTAGQuestion 4 (7 points)¶
In the previous questions, we have been specifying the DNA sequence by hand, but DNA sequences are usually long and are stored in files. A common file format is the FASTA format which looks similar to this:
>label1
ACTGTATCGATGCTAGCTACGTAGCTAGCTAGCTAGCTGACGTA
ACGATGTGCGAGGGTCATGGGACGCGAGCGAGTCTAGCACGATC
>label2
ACTGGGCTTGACTACGGCGGTATCTGACGGGCGAGCTGTACGAG
ACGGACTAGGGCGCGGCGGGGCGGATTTTCGAGTCGAGCGTTAT
The first line starts with a > which is immediately followed by a label (which might be the name of the gene for example). The sequence then starts on the second line, and may continue on several lines. It is common to limit the length of each line to 80, but this may vary from file to file. The sequence stops once either the file ends, or a line starts with >, which indicates that a new sequence is being given. There may be any number of sequences in a file.
Write a function read_fasta, that takes the name of a file (as a string) and returns a Python dictionary containing all the sequences from the file, with the keys in the dictionary corresponding to the label. If a sequence is given over several lines, you should remove any line returns and spaces. You should then be able to access the DNA for label1 with d['label1'] for example (if d is the name of the dictionary).
Use this function and the functions you have written above to read in the data/problem_1_question_4.fasta file and print out, for each sequence, the label, followed by the amino acid sequence (not the DNA sequence!).
Part 3 (5 points)¶
Question 5 (3 points)¶
Given several sequences with the same length, but which may include point mutations (i.e. individual nucleotides are changed), we want to try and find the most likely original sequence. For example, if we have the following sequences:
sequence 1: A C T C T
sequence 2: A C T C G
sequence 3: G C C C T
sequence 2: A C T C T
sequence 4: A T G C T
we can go through each position and find the most common nucleotide. To do this, we can first construct a matrix that looks like:
A: 4 0 0 0 0
C: 0 4 1 5 0
G: 1 0 1 0 1
T: 0 1 3 0 4
which indicates how many nucleotides of each type are found at each position, and from this we can see that the most common first base is A, the most common second base is C, and so on. The most common sequence is then ACTCT. This is the consensus sequence.
Write a function consensus_sequence that takes a dictionary of sequences (such as the one returned by read_fasta), and then returns the consensus sequence. Read the data/problem_1_question_5.fasta file using the function you wrote previsouly, and print out the corresponding consensus sequence. All the sequences in this file are the same length.
Note that the selection is based on the nucleotides!
Question 6 (2 points)¶
In some cases, it is useful to be able to identify the longest common sub-sequence between two sequences. For example, in the sequences ACTGCT and TGCCCT, the longest common sub-sequence is TGC (ACTGCT and TGCCCT). Note that these do not have to be at the same positon in each sequence.
Write a function, longest_common_sequence, that takes a dictionary of sequences (such as the one returned by read_fasta) and returns the longest common sub-sequence found in all the sequences.
Read the data/problem_1_question_6.fasta file, and print out the longest common sequence between all the sequences.